5.2 Segmentation: U-Net and its variants
This mainline page answers the Chapter 5 question "Why does segmentation work?" Full runnable scripts, demo outputs, and environment notes are collected in 5.6 Code Labs / Practice Appendix and
src/ch05/README_EN.md.
"U-Net is not just a network architecture, but a revolutionary thinking in medical image segmentation—proving that carefully designed architectures can surpass brute-force training on large datasets." — — Ronneberger et al., "U-Net: Convolutional Networks for Biomedical Image Segmentation", MICCAI 2015
Readers often feel classification is not enough in cases such as:
- measuring tumor volume;
- outlining the lung field precisely;
- defining where a radiotherapy target should stop slice by slice;
- describing the spatial relation between lesions and nearby anatomy.
⚡ U-Net's Success in Medical Imaging
Special Challenges of Medical Image Segmentation
Compared to natural image segmentation, medical image segmentation faces unique challenges:
| Challenge | Natural Image Segmentation | Medical Image Segmentation | U-Net's Solution |
|---|---|---|---|
| Data Scarcity | Millions of labeled images | Usually only hundreds | Skip connections enhance feature transfer |
| Boundary Precision Requirements | Relatively relaxed | Sub-pixel level precision requirements | Multi-scale feature fusion |
| Class Imbalance | Relatively balanced | Lesion regions usually very small | Deep supervision techniques |
| 3D Structure Understanding | Primarily 2D | Need 3D context information | Extended to 3D versions |
U-Net's Revolutionary Design Philosophy
U-Net's success stems from three core design principles:
- Encoder-Decoder Structure: Compress information like a funnel, then gradually restore
- Skip Connections: Directly transfer shallow features to avoid information loss
- Fully Convolutional Network: Adapt to input images of any size
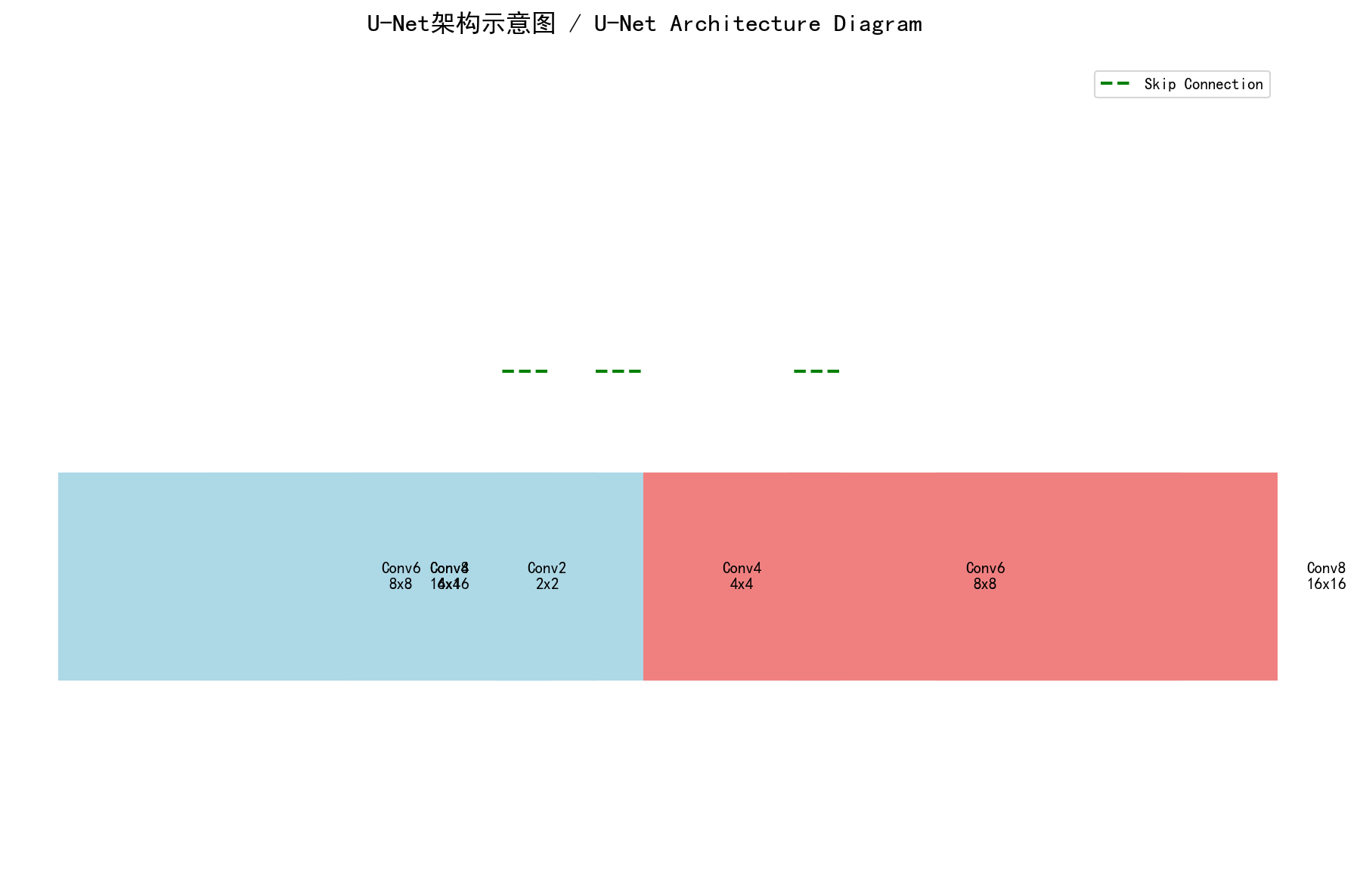 U-Net's core idea: Encoder extracts semantic features, decoder restores spatial resolution, skip connections ensure details aren't lost - Custom diagram
U-Net's core idea: Encoder extracts semantic features, decoder restores spatial resolution, skip connections ensure details aren't lost - Custom diagram
Intuitive explanation
Before thinking of segmentation as a complicated network, it is better to think of it as: turning image reading into measurable regions.
Classification answers whether something is present. Segmentation goes further and answers:
- where the region is;
- how large it is;
- what the volume is;
- how its boundary relates to nearby structures.
U-Net became classic not because it is simply deep, but because it captures one core tension:
- deep features are better at saying what something is;
- shallow features are better at preserving where it is.
So the network extracts semantics on the way down and reconnects spatial detail on the way back up.
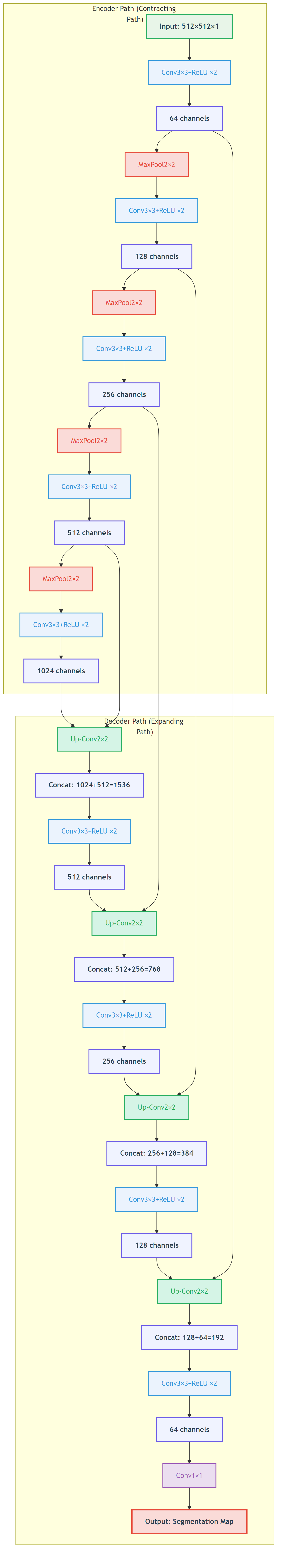 Figure: the key to U-Net is not just downsampling and upsampling, but the skip connections that bring boundary detail back.
Figure: the key to U-Net is not just downsampling and upsampling, but the skip connections that bring boundary detail back.
Core method
This section keeps only 4 key ideas.
1. Be clear about the output
Segmentation usually outputs not a single score, but a probability map or mask at the same scale as the input, or one that can be mapped back to the original image.
2. Preserve semantics and boundaries together
The encoder extracts high-level semantics; the decoder restores spatial resolution; skip connections prevent small structures and boundaries from disappearing completely during downsampling.
3. Keep labels perfectly aligned with images
In segmentation, one of the biggest failure modes is not weak augmentation but desynchronized transforms between image and mask. Once they drift apart, the model is no longer learning true boundaries.
4. Do not judge quality by loss alone
Dice, IoU, sensitivity, connected-component behavior, and post-processing results often say more about practical segmentation quality than training loss by itself.
Typical case
Case 1: Lung field segmentation
- Goal: obtain a lung mask from chest CT.
- Value: provides an ROI for later nodule detection, infection analysis, and quantitative measurement.
- Local code:
src/ch05/lung_segmentation_network/main.py.
Case 2: Organ or tumor segmentation
- Goal: precise contours for liver, pancreas, brain tumor, prostate, and similar targets.
- Value: volume estimation, disease burden tracking, and radiotherapy target delineation.
- Extensions: 2D U-Net, 3D U-Net, Attention U-Net, nnU-Net.
Case 3: Augmentation in segmentation tasks
- Goal: make image and mask undergo the same geometric transform.
- Local code:
src/ch05/medical_segmentation_augmentation/main.py. - Reminder: when the image changes, the label must change in sync.
Practice tips
The main text only keeps short code fragments for intuition; full training, evaluation, and visualization live in src/ch05/.
1. A minimal convolution block
class MultisequenceSegmentationUNet(nn.Module):
def __init__(self, num_sequences=4, num_classes=4):
super().__init__()
# Create independent encoders for each sequence
self.sequence_encoders = nn.ModuleList([
self.create_encoder(1, 64) for _ in range(num_sequences)
])
# Feature fusion module
self.feature_fusion = nn.Conv2d(64 * num_sequences, 64, 1)
# Shared decoder
self.decoder = self.create_decoder(64, num_classes)
def forward(self, sequences):
# Independent encoding for each sequence
encoded_features = []
for seq, encoder in zip(sequences, self.sequence_encoders):
encoded, skip = encoder(seq)
encoded_features.append(encoded)
# Feature fusion
fused_features = torch.cat(encoded_features, dim=1)
fused_features = self.feature_fusion(fused_features)
# Decode
return self.decoder(fused_features)Specialized Strategies for X-ray Image Segmentation
Anatomical Prior Constraints
class AnatomicallyConstrainedUNet(nn.Module):
def __init__(self, base_unet):
super().__init__()
self.base_unet = base_unet
self.anatomy_prior = AnatomicalPriorNet() # Anatomical prior network
def forward(self, x):
# Base segmentation result
segmentation = self.base_unet(x)
# Anatomical prior
anatomy_constraint = self.anatomy_prior(x)
# Constrained fusion
constrained_segmentation = segmentation * anatomy_constraint
return constrained_segmentation💡 Training Tips & Best Practices
Medical-Specific Data Augmentation Strategies
Medical image segmentation requires special consideration of anatomical and clinical constraints. Unlike natural image segmentation, medical augmentation must maintain anatomical reasonableness and clinical relevance.
Recommended Medical Augmentation Techniques
- Elastic Deformation: Simulate physiological motion (respiratory, cardiac movement) with non-rigid grid deformation
- Intensity Transformation: Simulate variation in different scanning parameters and protocols across institutions
- Noise Addition: Model real clinical environment noise to improve robustness to low-quality images from mobile devices or emergency scenarios
- Partial Occlusion: Simulate metal artifacts from implants or motion artifacts from patient movement
Augmentation Methods to Avoid
- Random Rotation: May destroy anatomical structure that should maintain fixed orientations
- Extreme Scaling: May introduce unrealistic deformations inconsistent with physiological changes
- Color/Hue Transformation: Medical images are typically grayscale with specific physical meanings (HU values, T1/T2 weightings)
Clinical Principle: All augmentation strategies must undergo physician verification to ensure they create clinically realistic variations rather than introducing medical artifacts.
Data Augmentation Strategies
Special data augmentation for medical image segmentation:
def medical_segmentation_augmentation(image, mask):
"""
Special data augmentation for medical image segmentation
"""
# 1. Elastic deformation (maintain anatomical reasonableness)
if np.random.rand() < 0.5:
image, mask = elastic_deformation(image, mask)
# 2. Rotation (multiples of 90 degrees)
if np.random.rand() < 0.3:
angle = np.random.choice([90, 180, 270])
image = rotate(image, angle)
mask = rotate(mask, angle)
# 3. Flip (left-right symmetry)
if np.random.rand() < 0.5:
image = np.fliplr(image)
mask = np.fliplr(mask)
# 4. Intensity transformation
if np.random.rand() < 0.3:
image = intensity_transform(image)
return image, mask📖 Complete Code Example: medical_segmentation_augmentation/ - Complete medical image segmentation augmentation implementation with multiple augmentation strategies and clinical validation
Medical Image Segmentation Augmentation Demonstration
Practical Enhancement Effect Display
To understand the impact of different augmentation techniques on medical image segmentation, we create a simulated CT lung image and demonstrate how four medical-specific augmentation techniques affect the image while maintaining clinical validity.
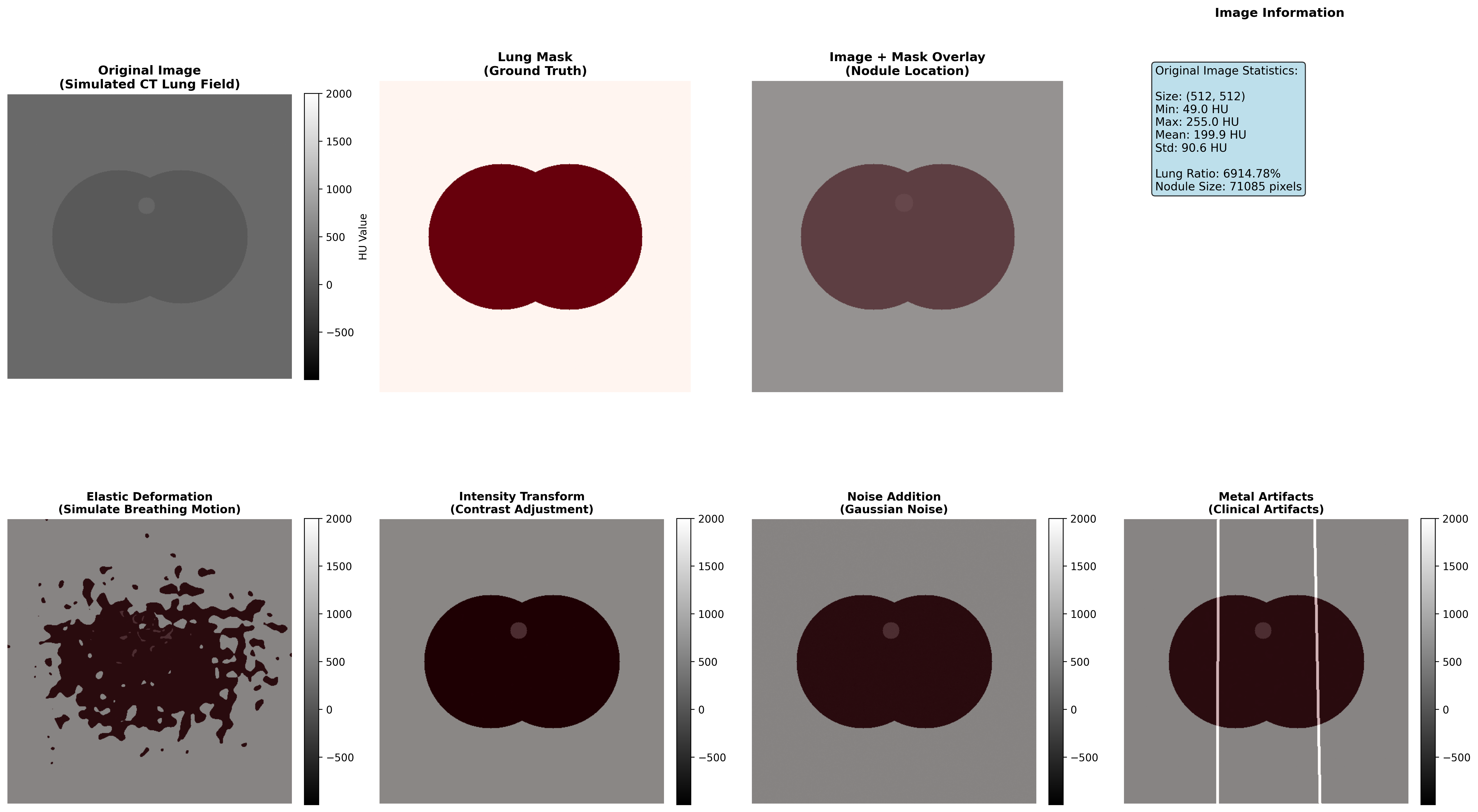
Demonstration of four medical-specific augmentation techniques: elastic deformation simulates respiratory motion, intensity transformation adapts to different scanning protocols, noise addition models real clinical environment, and metal artifacts simulate implant influences
Augmentation Technique Effectiveness Analysis
| Enhancement Type | Technical Principle | Clinical Significance | Application Scenarios | Implementation Caution |
|---|---|---|---|---|
| Elastic Deformation | Non-rigid grid deformation with interpolation | Simulate respiratory motion, cardiac pulsation | Thoracic and abdominal organs | Deformation intensity must remain within physiological range |
| Intensity Transformation | Contrast and brightness adjustment | Adapt to different scanning protocols, multi-center data unification | Multi-institution data fusion, cross-protocol analysis | Must preserve HU value medical meaning and tissue contrast relationships |
| Noise Addition | Gaussian or Poisson noise injection | Improve model robustness to low-quality images | Mobile devices, emergency scenarios, portable ultrasound | Noise characteristics should match actual device specifications |
| Metal Artifacts | Linear high-density streak simulation | Simulate metal implant influences (dental fillings, hip prostheses, pacemakers) | Orthopedic and dental imaging, cardiac device patients | Artifact morphology should match actual implant types |
Execution Results Analysis
Medical Image Segmentation Augmentation Analysis Report
def double_conv(in_channels, out_channels):
return nn.Sequential(
nn.Conv2d(in_channels, out_channels, 3, padding=1),
nn.ReLU(inplace=True),
nn.Conv2d(out_channels, out_channels, 3, padding=1),
nn.ReLU(inplace=True),
)2. The key skip-connection action: concatenation
import pydensecrf.densecrf as dcrf
class CRFPostProcessor:
def __init__(self, num_iterations=5):
self.num_iterations = num_iterations
def __call__(self, image, unary_probs):
"""
CRF post-processing: consider inter-pixel relationships
"""
h, w = image.shape[:2]
# Create CRF model
d = dcrf.DenseCRF2D(w, h, num_classes=unary_probs.shape[0])
# Set unary potentials
U = unary_probs.reshape((unary_probs.shape[0], -1))
d.setUnaryEnergy(U)
# Set binary potentials (inter-pixel relationships)
d.addPairwiseGaussian(sxy=3, compat=3)
d.addPairwiseBilateral(sxy=80, srgb=13, rgbim=image, compat=10)
# Inference
Q = d.inference(self.num_iterations)
return np.array(Q).reshape((unary_probs.shape[0], h, w))📈 Performance Evaluation & Model Comparison
Clinical Significance of Performance Metrics
Segmentation quality metrics have direct clinical implications:
| Metric | Clinical Application | Excellent Standard | Good Standard | Improvement Strategy |
|---|---|---|---|---|
| Dice Coefficient | Lesion volume assessment for treatment planning | >0.85 | >0.75 | Improve boundary accuracy through loss function refinement |
| Intersection over Union (IoU) | Overlap region calculation for surgical planning | >0.80 | >0.70 | Enhance overall consistency through architecture optimization |
| Sensitivity (Recall) | False negative control - avoiding missed lesions | >0.95 | >0.90 | Reduce false negatives through weighted loss or class rebalancing |
| Specificity | False positive control - avoiding over-segmentation | >0.90 | >0.85 | Reduce false positives through stronger anatomical constraints |
| Hausdorff Distance | Boundary deviation measurement for surgical navigation | <5 mm | <10 mm | Refine boundary precision through boundary-aware losses |
Clinical Decision Guidelines:
- For surgical planning: Require Dice >0.85 and Hausdorff <5mm
- For volume assessment: Require Dice >0.80 and IoU >0.75
- For lesion detection: Require Sensitivity >0.95 to minimize false negatives
Common Training Issues and Diagnostic Solutions
Issue 1: Model Over-predicts Background
Symptoms: Low Dice coefficient, high specificity Root Cause: Class imbalance in training data, excessive learning rate, insufficient data augmentation Solution Strategy:
- Adjust loss function weights: Increase lesion class weight by 2-5×
- Reduce learning rate by 50% and train longer
- Add aggressive data augmentation for minority class
Issue 2: Blurry Segmentation Boundaries
Symptoms: Low boundary Dice coefficient despite acceptable overall Dice Root Cause: Loss of fine spatial information in skip connections, insufficient decoder resolution Solution Strategy:
- Add explicit boundary loss term:
loss_total = loss_main + 0.3 × loss_boundary - Verify skip connection alignment between encoder and decoder
- Increase decoder intermediate channels
Issue 3: Complete Miss of Small Targets
Symptoms: Good performance on large lesions, completely missing small targets Root Cause: Large receptive field of deep layers, insufficient multi-scale information Solution Strategy:
- Implement multi-scale training: Train on multiple image resolutions
- Add dedicated small lesion branch in decoder
- Apply progressive training: Start with large targets, add small targets later
Evaluation Metrics
1. Dice Coefficient
Where:
: Predicted segmentation result : Ground truth annotation
2. Intersection over Union (IoU)
3. Hausdorff Distance
Hausdorff distance measures the maximum deviation of segmentation boundaries:
Where:
Performance Comparison of Different U-Net Variants
| Model | Dice Score | Parameter Count | Training Time | Applicable Scenarios |
|---|---|---|---|---|
| Original U-Net | 0.85-0.90 | ~31M | Moderate | 2D image segmentation |
| V-Net | 0.88-0.93 | ~48M | Longer | 3D volume data |
| U-Net++ | 0.87-0.92 | ~42M | Longer | Fine boundary requirements |
| Attention U-Net | 0.89-0.94 | ~35M | Moderate | Large background noise |
| nnU-Net | 0.91-0.96 | Variable | Auto-optimized | General scenarios |
🏥 Clinical Application Case Studies
Case 1: Brain Tumor Segmentation
Task Description
Use multi-sequence MRI to segment different brain tumor regions:
- Necrotic core
- Edema region
- Enhancing tumor
Data Characteristics
- Multi-modal input: T1, T1ce, T2, FLAIR
- 3D volume data
- Extremely imbalanced classes
U-Net Architecture Adaptation
class BrainTumorSegmentationNet(nn.Module):
def __init__(self):
super().__init__()
# Multi-sequence encoders
self.t1_encoder = EncoderBlock(1, 64)
self.t1ce_encoder = EncoderBlock(1, 64)
self.t2_encoder = EncoderBlock(1, 64)
self.flair_encoder = EncoderBlock(1, 64)
# Feature fusion
self.fusion_conv = nn.Conv2d(256, 64, 1)
# Decoder (4-class segmentation: background + 3 tumor classes)
self.decoder = UNetDecoder(64, 4)
def forward(self, t1, t1ce, t2, flair):
# Encode each sequence
_, t1_features = self.t1_encoder(t1)
_, t1ce_features = self.t1ce_encoder(t1ce)
_, t2_features = self.t2_encoder(t2)
_, flair_features = self.flair_encoder(flair)
def fuse_skip(upsampled, shallow_feature):
return torch.cat([upsampled, shallow_feature], dim=1)3. A minimal Dice implementation
import torch
def dice_score(pred_mask, true_mask, eps=1e-6):
inter = (pred_mask * true_mask).sum()
union = pred_mask.sum() + true_mask.sum()
return (2 * inter + eps) / (union + eps)🎯 Core Insights & Future Outlook
U-Net's Core Advantages:
- Skip connections solve deep learning feature loss problems
- Encoder-decoder structure balances semantic information and spatial precision
- End-to-end training simplifies the segmentation pipeline
Importance of Modality Adaptation:
- CT: Utilize HU value prior knowledge
- MRI: Multi-sequence information fusion
- X-ray: Anatomical prior constraints
Loss Function Design:
- Dice Loss addresses class imbalance
- Focal Loss focuses on hard samples
- Combined loss functions improve performance
Practical Tips:
- Data augmentation maintains anatomical reasonableness
- Multi-metric training process monitoring
- Post-processing improves final accuracy
Future Development Directions:
- Transformer-based segmentation models
- Self-supervised learning to reduce annotation dependency
- Cross-modal domain adaptation
🔗 Typical Medical Datasets and Paper URLs Related to This Chapter
Details
Datasets
| Dataset | Purpose | Official URL | License | Notes |
|---|---|---|---|---|
| BraTS | Brain Tumor Multi-sequence MRI Segmentation | https://www.med.upenn.edu/cbica/brats/ | Academic use free | Most authoritative brain tumor dataset |
| LUNA16 | Lung Nodule Detection and Segmentation | https://luna16.grand-challenge.org/ | Public | Standard lung nodule dataset |
| MSD | Multi-organ Segmentation | https://medicaldecathlon.grand-challenge.org/ | Public | Multi-organ segmentation challenge |
| ATLAS | Cardiac CT/MRI Segmentation | http://medicaldecathlon.grand-challenge.org/ | Academic use free | Cardiac segmentation dataset |
| KiTS21 | Kidney Tumor Segmentation | https://kits21.kits-challenge.org/ | Public | Kidney tumor segmentation |
| ISBI | Cell Segmentation | http://brainiac2.mit.edu/isbi/ | Public | Electron microscope cell segmentation |
Papers
| Paper Title | Keywords | Source | Notes |
|---|---|---|---|
| U-Net: Convolutional Networks for Biomedical Image Segmentation | U-Net segmentation network | arXiv:1505.04597 | Original U-Net paper, pioneering encoder-decoder architecture for medical image segmentation |
| U-Net++: A Nested U-Net Architecture for Medical Image Segmentation | Deep supervision segmentation | arXiv:1807.10165 | U-Net++ improvement, enhancing segmentation accuracy through deep supervision and dense connections |
| nnU-Net: A Framework for Automatic, Deep Learning-Based Biomedical Image Segmentation | Automatic segmentation framework | Nat Methods 18, 203–211 (2021) | nnU-Net automation framework, achieving SOTA performance in multiple medical segmentation tasks |
| V-Net: Fully Convolutional Neural Networks for Volumetric Medical Image Segmentation | 3D medical segmentation | 2016 Fourth International Conference on 3D Vision | V-Net, fully convolutional network designed specifically for 3D medical image segmentation |
| Attention U-Net: Learning Where to Look for a Pancreas | Attention mechanism segmentation | arXiv | Introducing attention mechanism in medical segmentation to improve target region recognition |
| Deep Learning for Brain Tumor Segmentation: A Survey | Brain tumor segmentation review | Springer Journal: Complex & Intelligent Systems | Comprehensive review of deep learning methods and comparisons for brain tumor segmentation |
| 3D U-Net: Learning Dense Volumetric Segmentation from Sparse Annotation | 3D sparse segmentation | arXiv:1606.06650 | 3D U-Net extension, suitable for sparsely annotated 3D medical image segmentation |
Open Source Libraries
| Library | Function | GitHub/Website | Purpose |
|---|---|---|---|
| TorchIO | Medical Image Transformation Library | https://torchio.readthedocs.io/ | Medical image data augmentation |
| nnU-Net | Automatic Segmentation Framework | https://github.com/MIC-DKFZ/nnunet | Medical image segmentation framework |
| MONAI | Deep Learning Medical AI | https://monai.io/ | Medical imaging deep learning |
| SimpleITK | Medical Image Processing | https://www.simpleitk.org/ | Image processing toolkit |
| ANTsPy | Image Registration and Analysis | https://github.com/ANTsX/ANTsPy | Advanced image analysis |
| medpy | Medical Image Processing | https://github.com/loli/medpy | Medical imaging algorithm library |
| DeepLabv3+ | Semantic Segmentation | https://github.com/tensorflow/models/tree/master/research/deeplab | DeepLabv3+ implementation |
The next section moves to classification and detection because not every workflow needs a fine contour first. Many clinical pipelines start by asking: is there disease, what is it, and roughly where should we look?